Every engineering discipline requires and eventually develops its own specialized standards for communicating information about the structure and function of designs. This June, the synthetic biology community took another important step toward effective sharing of designs, as ACS Synthetic Biology became the first scientific journal to officially adopt standards for how genetic constructs should be depicted and how the designs of engineered organisms should be recorded and shared. In particular, ACS Synthetic Biology has chosen to adopt the two current Synthetic Biology Open Language (SBOL) standards: SBOL visual for drawing genetic diagrams, and the SBOL data standard for documenting the design of engineered genetic constructs.In a viewpoint article in the June issue of ACS Synthetic Biology, managing editor Ranjini Prithviraj joined with members of the SBOL community to announce the journal’s new policies. The journal was motivated to adopt standards by the challenges faced by its authors, as editor-in-chief Chris Voigt is quoted in the article: “As the complexity of synthetic biology projects grow, it will be critical to standardize the exchange of designs and data.” SBOL fits the bill: it is a free and open standard, developed by a broad international community and already in use by many synthetic biologists both in academia and industry.Now that the journal has officially adopted SBOL, its author guidelines have been adjusted to recommend that genetic constructs should be depicted using SBOL Visual and that SBOL 2 is the preferred format for nucleic acid sequences. SBOL visual provides a palette of symbols for making clear and precise diagrams of genetic designs; these will mostly be quite familiar and easy for authors to adopt, as SBOL visual was developed mainly by organizing and distilling a large number of prior published genetic diagrams in order to obtain a single coherent system.For the SBOL data standard, the journal has also incorporated an SBOL-supporting data repository into its reviewing process. Authors submit their supplementary genetic design information to https://acs-registry.jbei.org, which validates the constructs (and can also convert FASTA and GenBank into SBOL). During the reviewing process, designs are kept private and shared only with the paper’s editor and reviewers. If the article is then accepted, the designs are automatically made public and linked to the article, while if it is rejected they remain private to the author.
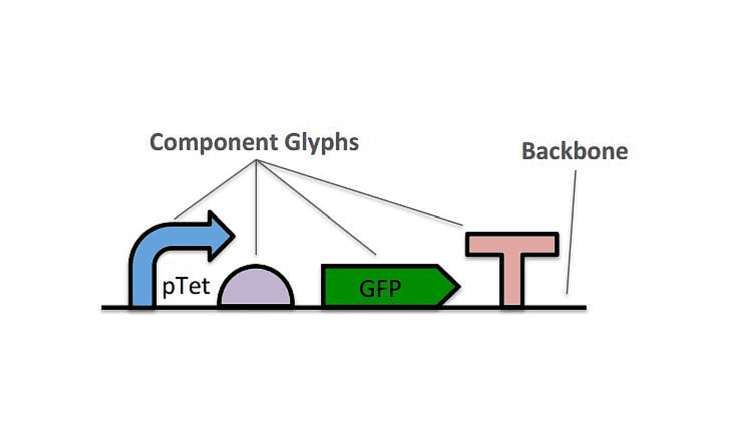
Genetic construct diagrammed with SBOL visual.The SBOL data standard differs from prior standards, like the GenBank file format, in that it has been designed to serve the specific needs of organism engineering. In addition to documenting nucleotide sequences and sequence features, the SBOL 2 data standard also allows synthetic biologists to incorporate other design elements, such as proteins and metabolites, to describe the way in which a design is organized from smaller components, how the structure of a design relates to its expected functional interactions, such as gene expression and regulation or the processing of metabolites by enzymes. All of this information can then be linked to external resources, such as computational models, characterization data, or protocol specifications, allowing a single SBOL design to serve as a “hub” integrating all of the available information about a design.As SBOL becomes more widely adopted, synthetic biologists should expect to find a growing body of well-documented and reproducible synthetic biology systems, as well as to contribute to this body themselves. As has been the case in every other field of engineering, this should make engineering organisms faster and cheaper, allow more complex systems to be engineered, and lower the barrier to entry by new practitioners. “Effective engineering demands clear communication and the ability to integrate components from many sources,” writes SBOL Steering Committee member Jacob Beal, “Safe and ethical engineering practices demand them even more so.” This will be accompanied by the emergence and maturation of standards in related areas like protocol automation, calibration of measurements, and characterization of strains and devices, further enhancing the acceleration of synthetic biology engineering.From its beginnings eight years ago at a meeting in Seattle in 2008, SBOL has been continuously developed by an active international community, with the first versions of both the visual and data standards released in 2009. The community now numbers over 100 members who hail from more than a dozen countries. The current version of SBOL visual was released in 2013, while SBOL 2 was ratified and adopted by the community more recently in 2015, and both the SBOL standards and a wide variety of SBOL-supporting computational tools continue to be refined in response to the evolving needs of the synthetic biology community. More information about SBOL can be found at sbolstandard.org, including resources for using SBOL, a list of SBOL-supporting tools, and information on how to contribute.
Read More
Newletter & More

SynBioBeta
Join the innovators shaping the future with SynBio + AI. From health to ag, materials & more—be part of the revolution.


















