Measure, pipette, mix, repeat. No doubt performing experiments in biotechnology are mundane tasks. In addition, the samples being handled are often in microscopic amounts and sensitive to the slightest of variations, therefore the experiments are difficult to reproduce. Most researchers would agree, such tasks are better left to robots, which also buys them more time to pursue the intellectual part of their jobsThat’s where automation platform companies like Synthace, Transcriptic, and Emerald Cloud fill in. Their goal is to make biology easier to engineer and outcomes more predictable. Organism engineering capabilities of such organism companies often suit multiple industry sectors. Synthace relies on an in-house programming language for the purpose and recently announced its release to the global community in SynBioBeta SF conference held in November.
Antha and Its Workflows
Antha (@Anthalang) can, in simple words, be described as a programming language for experiments in biology, automated or otherwise. This high-level language allows users to develop, execute, and analyse biological experiments faster. The language is open source and allows for dissemination of protocols to the wider scientific community.Antha simplifies protocols into reusable components that can be exchanged or removed to produce simple, reproducible, and scalable workflows (henceforth used to describe a process or series of processes). It systematizes not just the parts, but the whole processes. Thus, it introduces two components of engineering generally lacking in the engineering of biology -- standardization and modularity.
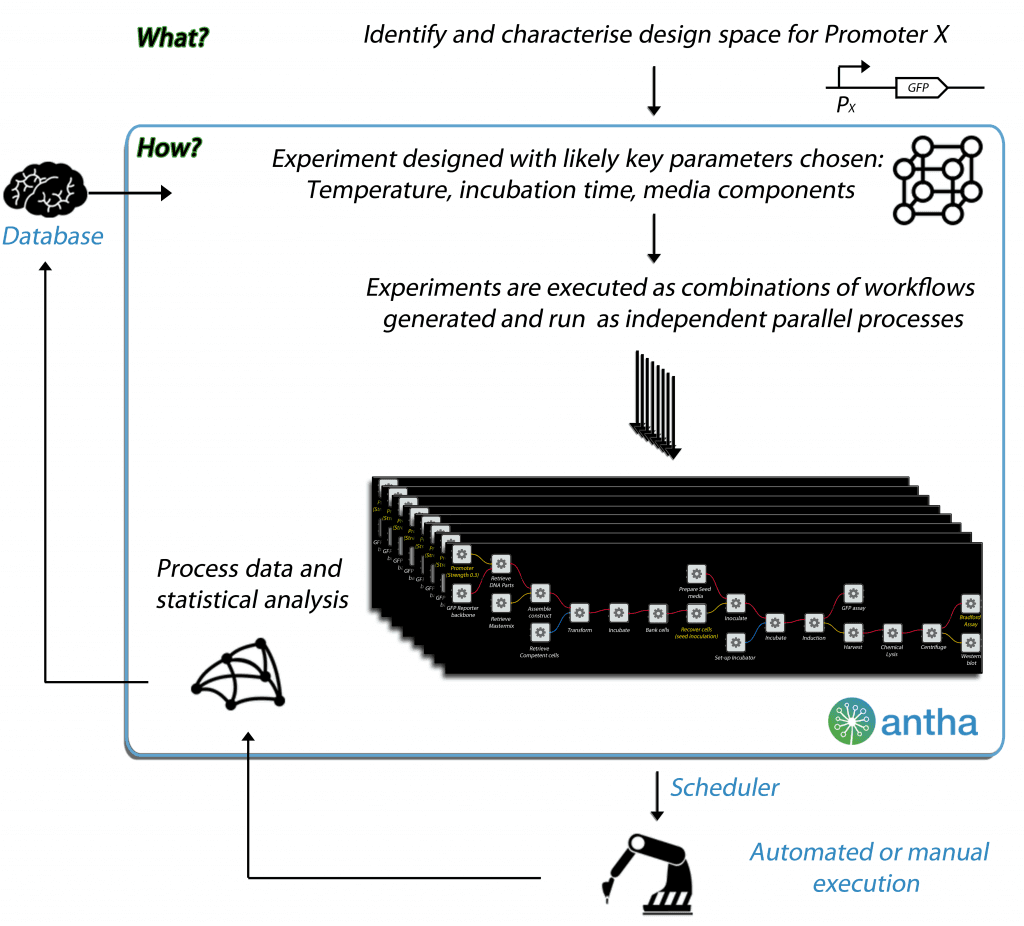
In addition to being a programming language, it manages the laboratory information and experimental data, and serves as a quality assurance system. In a conventional setting, only a few parameters are recorded while many other parameters might be affecting the outcome in various degrees. With Antha’s robust system, a multitude of variables that would usually go unnoticed are accounted for.As a result, as long as the sequence of workflows is the same, the differences in the lab equipment and the experimenter will be accounted for in the results. The visual representation can be downloaded for use in the article.
It Is Beautiful
It utilizes a collection of easily comprehensible blocks. These include imports, parameters, data, inputs, outputs, requirements, setup, steps, analysis, and validation. Imports refer to element names, additional elements if any, and the code needed for an execution. Setup refers to the task that must be performed the first time an element is subjected to a workflow. The inputs and outputs are the reagents and labware used and the products formed respectively. The rest mean the same things they do normally.The syntax of Antha is a derivative of the Go language from Google. Any workflow is executed as an array of parallel processes and described from the standpoint of the smallest appropriate unit of reaction. In terms familiar to the conventional biologist, a set of actions in a protocol are performed with the most influencing factor in check.Here’s the Antha syntax to perform what is probably one of the most repeated biology experiment in history, the Bradford’s test to estimate proteins.// Input parameters for this protocol (data)
Parameters {
var SampleVolume Volume = 15.(uL)
var BradfordVolume Volume = 5.(uL)
var Wavelength Wavelength = 595.(nm)
var ControlCurvePoints uint32 = 7
var ControlCurveDilutionFactor uint32 = 2
var ReplicateCount uint32 = 1 // Note: 1 replicate means experiment is in duplicate, etc.
}All the parameters of the experiment such as the volumes used and the wavelength at which absorbance is recorded are stored as variables for use in workflows later.Validation {
if SampleAbsorbance > 1 {
panic("Sample likely needs further dilution")
}
if (RSquared < 0.9) {
warn("Low r_squared on standard curve")
}
if (RSquared < 0.7) {
panic("Bad r_squared on standard curve")
}
// TODO: add test of replicate variance
}Bradford test works only within a narrow range of protein concentration. If the sample contained too much or too low protein, the test needs to be repeated. The sample cannot be recovered, but the wastage could have been avoided by a non-destructive validation check. In more sophisticated tests, only a fraction of the sample is used up in destructive validation. Another advantage.If you are interested in programming, you can check other blocks of this particular example at Antha’s website or browse through its repositories on Github.
Why Coding Biology Experiments Matters
Sean Ward, founder-CTO of Synthace best sums up the purpose of Antha in Synthace’s announcement of Antha:“The critical thing is that once one person has made a biological process work and encoded it in Antha, it’s then a problem that is solved for everyone, for ever” he says, “and achieving the aspirations of synthetic biology is only possible by sitting atop a foundation which allows everyone to grow and disseminate new ways of working”.Automation is improving reproducibility by allowing exact control of the parameters. But, it is possible that the irreproducibility is more due to partial publication of results than experimental variability. Automation could still help by allowing large number of replicates to be run, but the nature of the experiments need to change.It is required that all failed and successful experiments are recorded, accurate to every minor detail, to make any sense of the chaos inherent of biological systems. This is important if synthetic biology is to go from being quite a bit predictable to very predictable. It would be a paradigm leap in biological experimentation.
Read More
Newletter & More

SynBioBeta
Join the innovators shaping the future with SynBio + AI. From health to ag, materials & more—be part of the revolution.
















