Dow AgroSciences and TeselaGen Biotechnologies To Partner in Developing Biological Design Platform
Dow AgroSciences sees a big opportunity in natural products. That chiefly explains a half-dozen collaborations the Indianapolis-based maker of crop protection and plant biotechnology solutions has announced in recent months, including an initiative unveiled today with San Francisco-based TeselaGen Biotechnology to build a “state-of-the-art biological design platform.”
Dow AgroSciences and TeselaGen are building “AutoCAD for biology
“It’s part of our overall strategy to find winners,” says Steven Webb, R&D director at Dow AgroSciences. “We have a lot of smart people but we also know there are many more out there in adjacent spaces. We’re spending a lot of time looking on the outside looking for exciting opportunities to leverage their experience and accelerate our effort to find new solutions.”As an example of a recent innovation in the natural products space, Webb points at a new fungicide, Inatreq, that Dow released last year in the battle against Septoria tritici in cereal crops, a problem that’s especially serious in Europe.The new technology that Dow AgroSciences and TeselaGen are using to build this new platform can be described as “AutoCAD for biology,” says TeselaGen co-founder and CEO Mike Fero. “It’s the only cloud-based platform on the market that automatically generates cost-optimized DNA construction protocols.” A patent on the technology is pending. Fero said he expects it to be awarded within 30 days.DNA construction is the process by which multiple genes or fragments of DNA sequences are assembled. It incorporates DNA sequence fragments from a variety of organisms into a self-replicating genetic element, such as a bacterial plasmid, that will propagate the assembled parts in a host cell.Based on the j5 algorithm that Nathan Hillson developed five years ago at the Joint BioEnergy Institute and licensed from Lawrence Berkeley National Laboratory by the TeselaGen team of Hillson, Fero and Eduardo Abeliuk, the platform features a graphical interface that enables users to design a DNA construct or combinatorial libraries through the arrangement of individual part icons that abstractly represent underlying DNA sequences. Easy-to-understand outputs detail the resulting designed experimental protocols, providing instructions that can either be followed by a person in the laboratory or fed directly into a robotic platform for a machine to carry out.
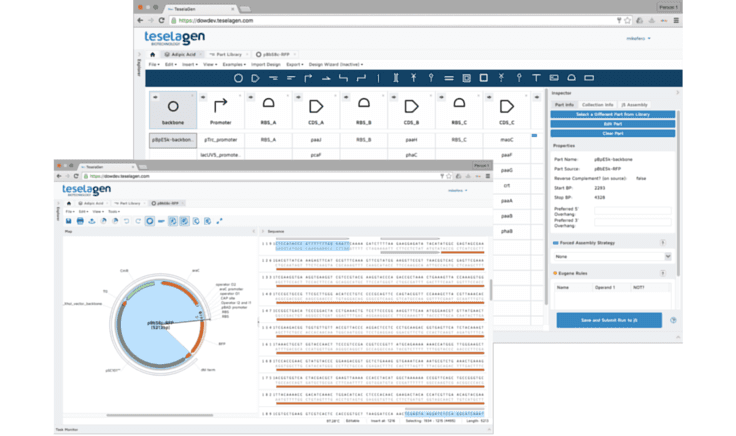
Embedded in the TeselaGen platform, the j5 algorithm can quickly determine the optimal flanking sequences that should be attached to each DNA part to produce the desired recombinant DNA at the least expense, in a way that can be executed either by hand or by robots.“Researchers can focus on investigating their primary interests, rather than preparing the DNA that’s merely a tool in their experiments,” Fero says.“The more complex the task, the more time and money the program can save,” Fero says, noting that the greatest savings are found in combinatorial libraries, collections of hundreds to hundreds of thousands of related DNA assemblies, each with a different combination of genes or parts that perform similar functions in different organisms. Combinatorial libraries can be screened to identify the gene combination that, when transferred into a desirable host organism, results in the most productive enzyme pathway.The three founders met at Stanford while pursuing advanced studies in systems biology and genomics. In developing the TeselaGen platform, they’re continuing to receive support from two National Science Foundation SBIR (Small Business Innovation Research) grants. Dow AgroSciences announced a similar partnership with Radiant Genomics in late February. As in its new collaboration with TeselaGen, the goal is to slash the time required to identify new lead molecules from multiple years to mere months. “It’s all about our ability to manage knowledge more quickly to do the design, build, test and learn cycle faster and faster,” says Webb.


















